0
You have 0 items in your cart
We will be closed Monday, September 2, 2024, in observance of Labor Day. Normal business hours will resume on Tuesday, September 3, 2024! Don't forget - our drop boxes are always open!
PDF of information below: Soil Enzyme Information Rev 1.0
Soil macro and microbiological activity is responsible for storing, transforming and cycling nutrients in the soil. Soil macrobiology, such as arthropods and worms, create large tunnels in the soil for air and water to enter the soil, mix soil by bringing organic matter to the soil surface and partially decompose organic residues. Microbiology, such as bacteria, fungi, and actinomycetes (collectively referred to as microbes), finish decomposing organic residues, complete nutrient cycles, improve aggregation and build soil organic matter (SOM). In order to transform and release nutrients in the soil, soil microbes depend on an extensive variety of enzymes to complete this process.
Most microbes in the soil exist under starvation conditions. Fierce competition for food, water and space within the soil causes many microbes to live in a dormant state until resources become available. Often, soil microbes will gather around nutrient rich areas that contain ideal conditions (i.e. water, temperature, pH) for nutrient uptake. These locations, known as microsites, are prominent in the rhizosphere where gatherings of soil microbiology, collectively referred to as soil microbial communities, can be found attached to root hairs, on plant residues and on and within aggregates. Soil microbial communities cluster around living roots because of the unique relationship microbial communities have with plant roots.
Plants are dependent on roots to anchor themselves in the soil and allow nutrient and water uptake. In addition to stabilizing the plant, roots also secrete a vast array of unique chemical compounds that attract and repel certain soil microbes, build soil aggregates, change the soil pH and inhibit the growth of competing plants. These secreted chemicals, referred to as root exudates, help cultivate the microbiome that exists around the root and is exclusive to soil type, climate and vegetation of the area. Since the root has limited exposure to necessary nutrients for growth and production, root exudates are released in the rhizosphere to attract microbes to the root area. The release of simple nutrients to support soil microbial communities is exchanged for microbial scavenging of essential plant nutrients. Like plants, soil microbes require macro and micro nutrients to survive, reproduce and perform basic continuous maintenance. The soil nutrients in direct contact with the plant root are limited, requiring soil microbes to rely on various chemical and biological processes to obtain nutrients from the soil environment.
Most nutrients in the soil are locked away in large, complex compounds that soil microbes are unable to pass through their cell wall. In order to breakdown these large, complex compounds, soil microbes produce specific protein catalysts, known as enzymes, to fracture complex compounds into simpler forms of nutrients. Thus, soil enzymes are important tools for soil microbes to create accessible nutrient forms that are small enough for both the plant and the microbe to pass through their cell wall.
Soil nutrient availability varies over space and time. Often, soil nutrient supply does not align with plant and microbial nutritional demand. If the nutrient supply is sufficient to meet plant and microbe demand, little to no production of nutrient specific enzymes are created. If the nutrient supply cannot be met, enzymes are produced to fill the nutrient gap. The production of enzymes is costly to a microbe’s limited resources; thus, enzyme production is mostly carried out when the need is great, and the substrate is available. The abundance, availability and activity of enzymes is not only dependent on production but also on the location of soil enzymes in the soil environment.
Large complex compounds from living and dead plants, animals and microbes are often physically and chemically attached to clays, minerals or other organic compounds in the soil. These soil components shield important sources of nutrients from microbial access; requiring enzymes to not only be present but also mobile. Enzymes exist predominantly in three different forms to access nutrients: intracellular, free extracellular and adsorbed extracellular (See Fig 1).
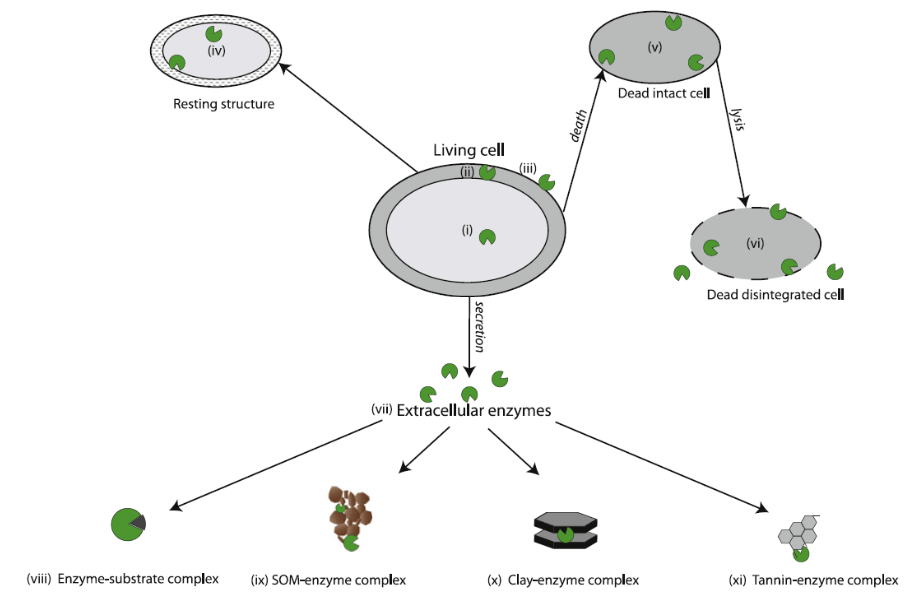
Figure 1: Enzyme locations within the soil as described by Burns 2013
All soil enzymes are created within the microbial cell. Some enzymes, known as intracellular enzymes, exist primarily within the cell or within the cytoplasm of the cell and act on substrates that are small enough to pass through the cell but require additional processing prior to being used by the cell. These enzymes can also prepare other proteins and enzymes for the hostile soil environmental. Intracellular enzymes can exist only within cell and upon death and, can be released into the soil environment through cell lysis (the rupturing of a cell). These enzymes are quickly denatured or attacked by proteases, an enzyme designed to breakdown a wide variety of proteins. Extracellular enzymes, or enzymes designed to exist outside the cell, can become attached to the cell wall. If a cell develops a polysaccharidic coat or biofilm, the cell may create a scaffold-like structure, known as a cellulosome, which protrudes attached soil enzymes into the surrounding soil environment. These enzymes can orient their active sites to stay open and can protect areas of the cell that may be subjected to attack by predators. Other extracellular enzymes are designed to enter the soil environment. By adjusting the structure of the enzymes, these extracellular enzymes are better able to withstand environmental stressors, such as temperature and pH, that cause denaturing and have better resistance to predation than intracellular enzymes. Once exuded from the parent cell, these enzymes, known as free extracellular enzymes, exist within the soil solution and are an important mode of enzyme transportation in the soil environment. Through diffusion, these enzymes spread throughout the soil environment, but often become stabilized by becoming adsorbed (attached to the surface of the material) or absorbed (attached within the material) with clay minerals, soil organic matter or humic material. Once an enzyme becomes stabilized in the soil, these enzymes are referred to as extracellular enzymes. If the active site of an extracellular enzyme is still exposed to the soil environment, the adsorbed enzyme can continue breaking down substrates into plant and microbial available nutrients long after the parent microbial cell has died. Adsorbed enzymes can also be protected from predation and other denaturing situations. Absorbed enzymes are often not exposed to substrates and become ineffective, unless the enzyme becomes exposed to the soil environment. The ability of extracellular enzymes to continue acting on substrates within the soil environment may be an important source of response to changes in substrate availability in soil that acts as an early alert system to microbial communities. This could allow communities to become more efficient in enzyme production or, if enzymes are active enough, the microbe may conserve those resources required to create enzymes by not producing them.
PDF of this information: Soil Enzyme Information Rev 1.0
Each enzyme is constructed by microbes using long chains of amino acids that are distinctively folded to form active sites specific to select substrates. Substrates, or the complex compound that will be acted upon by the enzyme, bind to the enzyme on the active sites to form an enzyme-substrate complex. At this point, depending on the type of enzyme, substrates can either become fused together to create a new product or bonds within the substrate are fractured and separate products will be released. This process is often viewed as a “lock and key” mechanism (See Figure 2) where the shape of the substrate must match the shape of the active site in the enzyme. After this process is completed, the enzyme is ready to accept another substrate. If the active site of an enzyme becomes permanently altered due to extreme changes in environmental conditions (i.e. pH and temperature), the substrate can no longer bind to enzyme and no products will be created. This process is known as denaturing and can not be reversed.
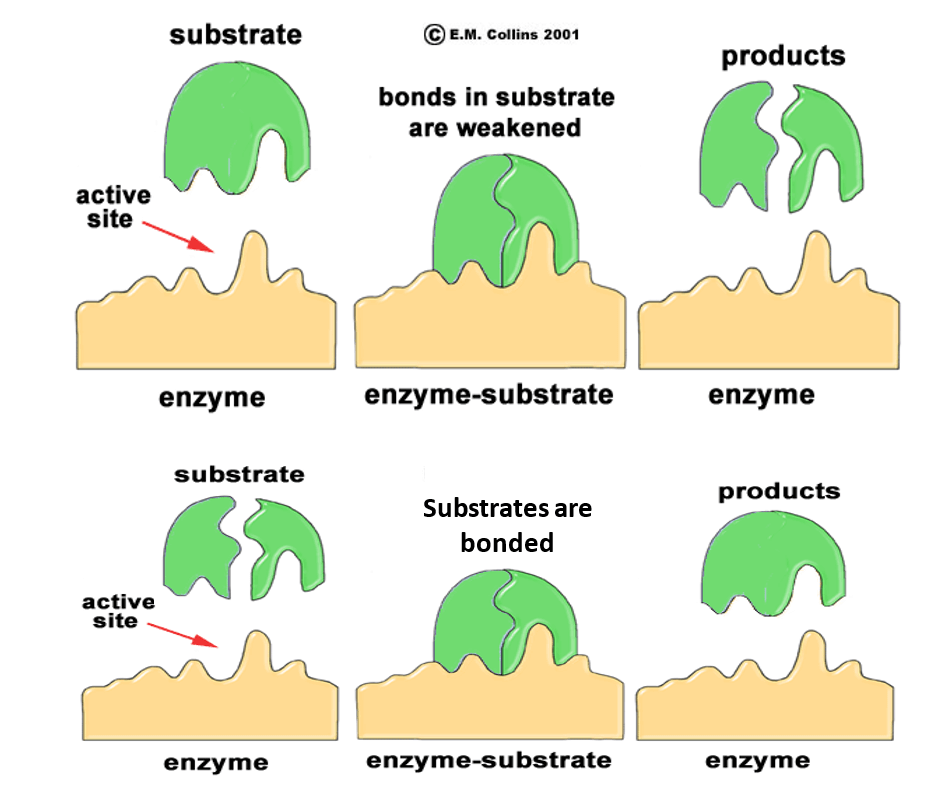

Figure 2: The “Lock and Key” Enzyme Mechanisms
(Image from E.M. Collins 2001)
The particular structure of soil enzymes allows only one kind of substrate to be processed and the activity of many enzymes is required to completely breakdown a complex compound. Soil enzymes target specific complex compounds such as proteins, carbohydrates, amino sugars, organic phosphates, and lignins to release readily available nutrients in the soil. For instance, cellulose is a basic structural component of plant cell walls that is composed of a long chain of bonded sugar molecules. These sugar molecules, predominantly made of glucose, are an important source of energy for soil microbes. Soil microbes produce a collective group of soil enzymes, known as cellulases, to disassemble the ridgid structure of cellulose to release glucose molecules. The cellulose structure is fractured by three dominant groups of enzymes: endoglucanases, exoglucanases and β-glucosidases (See Figure 3). Endoglucanases randomly cleave the large chained cellulose into smaller fractions while exoglucanases further reduce the cellulose chain to cellulobiose units (two linked glucose molecules). Finally, β-glucosidases cleave the cellulobiose molecules to release glucose as a readily available nutrient for microbe uptake. β-glucosidase activity is a very commonly monitored enzyme because it is not only responsible for the energy flow in the soil system but is only produced when cellulobiose is abundant.
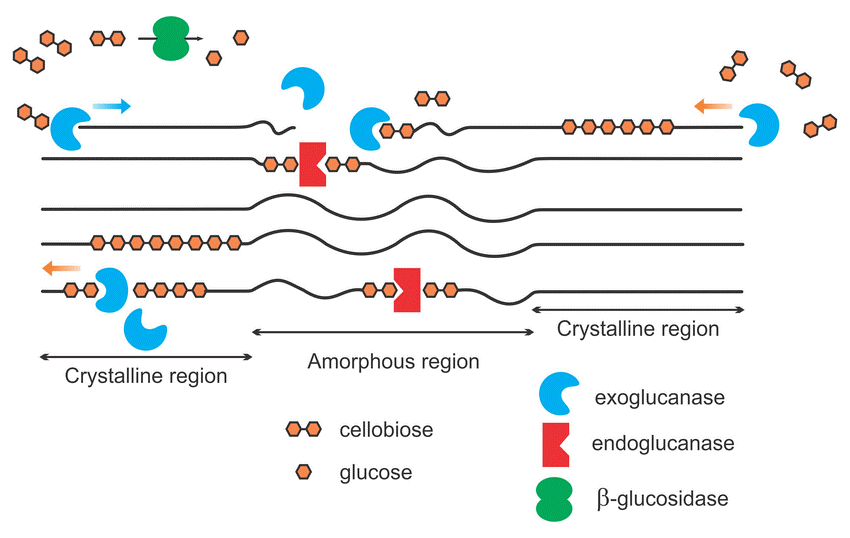
Figure 3: Simplified schematic of cellulase enzymes acting on cellulose
The long-chained cellulose is fractured by the accumulated effort of exoglucanases, endoglucanases and β-glucosidase enzymes to release glucose in the soil environment. (Image from Akhtar 2016)
Understanding the role of soil enzymes and monitoring the activity of specific enzymes provides earlier indication of how soil nutrient cycles are changing or responding to any changes in nutrient or energy supply and demand within the soil enviornment. Thus, soil enzymes are often measured due to their rapid response to soil management practices and are often referred to as “bio-indicators” of soil health.
The creation and distribution of enzymes reflects the availability of substrates in the soil environment. The specificity of soil enzymes provides a unique insight into early changes in the specific cycles in the soil environment. Although several enzymes are involved in nutrient and organic matter cycling, specific enzymes have been identified due to their vital role in soil metabolic processes, sensitivity to changes in soil management and ease of measurement. By monitoring select enzyme activities, the impact of soil management decisions on important cycles in the soil may be detected earlier than many other soil measurements.
PDF of this information: Soil Enzyme Information Rev 1.0
Soil enzyme testing at Ward Laboratories, Inc. is conducted by analyzing the consumption of a substrate and the release of a colored product. The consumption of product is measured over time and results are expressed as a rate of enzyme activity. By using controlled soil conditions (e.g. pH, temperature), the enzyme activity rates can indicate the potential activity for the soil enzymes under ideal conditions. This allows a comparison of potential enzyme activities between different soil management practices. Similarly, the same site can be tracked over time to monitor subtle changes in microbial dynamics and provide an indication of the microbial community response to changing soil environmental conditions and management.
Soil enzyme activity is reported as a rate of product (or color) produced (i.e. β-glucosidase is reported as ppm pNP g-1 soil h -1). Interpretation of soil enzyme activity requires an understanding of nutrient or organic matter cycling. Often, healthy, active systems have increased enzyme activity, relating to better cycling of nutrients and organic matter quality in the soil. Nevertheless, sites with recent disturbances may have higher activity levels due to increased substrate availability when compared to non-disturbed sites. For example, a conventional tillage field versus a no-till field in its first year of transition may indicate the tilled field has higher enzyme activity. This is because the act of tilling a field provides aeration and better distribution of substrates to microbes. The increased activity cannot be fully sustained in this system and often causes the enzymes to access the nutrients in organic matter, leading to a loss of organic matter the following year. A list of common soil characteristics and soil management impacts can be found in the interpretation guide.
Additional information will be added to the website as new information becomes available. Any questions regarding soil enzyme testing may be directed to Ward Laboratories, Inc at biotesting@wardlab.com.
Sampling for soil enzymes testing is slightly different than regular soil testing. It is very important to treat all the samples the same. Listed here are general guidelines to sample for all Soil Enzyme Tests.
Additional information will be added to the website as new information becomes available. Any questions regarding soil enzyme testing may be directed to Ward Laboratories, Inc at biotesting@wardlab.com.
Active Site: the section of an enzyme that acts on the substrate to release nutrients.
Aerobic microbes: microbes that require oxygen for energy, growth, reproduction and cellular respiration.
Absorption: molecules or ions that permeate a material such as a soil particle.
Adsorption: adhesion of a molecule or ion onto the surface of a solid such as a soil particle surface.
Aggregates: a grouping of soil material that is loosely formed by chemical and biological activity. The size and distribution of aggregates contributes to soil structure and allows air and water flow within the soil.
Anaerobic Microbes: microbes that require oxygen poor environments for energy, growth, reproduction and cellular respiration.
β-glucosidase (BG): a carbon cycling enzyme responsible for the release of glucose into the soil environment.
Bioindicator: a measurable biological presence or action that can be used to indicate the soil ecosystem’s status or “health”.
Bulk pH: an average measurement of hydrogen ions (H+) in soil solution.
Catalyst: presence of a substance that causes an increase in a reaction without being physically altered.
Cell Lysis: the rupture of a cell wall or membrane.
Cellulase: group of enzymes responsible for the breakdown of cellulose.
Cellulose: long-chained glucose moleculesthat serve as main structural components in the cell wall of plants.
Cellulosome: scaffold-like structure made up of discrete multi-enzyme complexes that protrudes from the surface of the cell wall and into the soil environment to aid in exposing enzymes to substrates.
Cofactor: an element or compound that must be present for an enzyme to function.
Complex Compounds: a broad term used to indicate a grouping of simpler substances (compounds or ions) that are connected by chemical bonds.
Enzyme: a protein catalyst created by animals, plants and soil microbiology to breakdown complex compounds into simpler molecules.
Enzyme Activity: the action of a protein catalyst on a substrate in the soil. Enzyme activity is expressed as the released product per gram of soil per hour.
Extracellular Enzymes: enzymes that exist outside the parent cell. Extracellular enzymes can refer to enzymes within the soil solution or adsorbed to the surface of organic matter, clay particles or other soil complexes. Extracellular enzyme activity is a large contributor to overall enzyme activity.
Exudate: a substance or moleculereleased into the rhizosphere to colonize bacteria and fungi.
Free Extracellular Enzymes: enzymes that exist outside the parent cell and within the soil solution.
Decomposition: the breakdown of a material to release nutrients into the soil.
Degradation: a breakdown of a compound into simpler forms (ions or compounds).
Denature: a permanent, physical alteration of the enzyme structure that prevents the enzyme from further acting on any substrates.
Dissolution: dissolving a soil solid.
Functional Diversity: a term used to describe a community based on the variety of biological processes, traits or characteristics of a particular ecosystem.
Glucose: a simple sugar that acts as an easy source of energy for soil microbes.
Immobilize Enzyme: an enzyme that is attached to an inert, insoluble material.
Intracellular Enzymes: enzymes that are created and reside within the microbial cell.
Inorganic Fertilizers: a chemical that releases plant available forms of nutrients quickly (e.g. ammonium nitrate).
Microbes: a broad term used to reference soil microorganisms such as bacteria, fungi and actinomycetes that require a microscope to see.
Microsite: a small section of soil that often contains ideal environmental conditions and rich resources such as nutrients, water, temperature, and pH for microbial proliferation. These areas are often contained in the rhizosphere around roots, within plant residues or on aggregate surfaces.
Microsite pH: localized site in the soil, such as on the surface of a soil particle or organic matter, where high chemical or biological activity has caused a change in pH. This pH often differs from bulk pH.
Mineralization: the conversion of inorganic forms of nutrients to organic forms of nutrients through microbial action.
Organic Fertilizers: sources of soil nutrients that originate from animal matter, manure and vegetation (e.g. applying manure to a field).
Organic Amendments: material added to the soil that originates from plants or animals.
Phosphatase: a group of enzymes that are responsible for the cycling of organic phosphates.
Phosphodiesterase (PHD): a soil enzyme responsible for the degradation of nucleic acids, phospholipids and other diesters which constitute a main component of fresh organic P inputs in soil.
Phosphodiesters: bonds that link deoxyribose to phosphorusand make up the backbone of nucleic acid.
Residue: undegraded material that has been left in a field (e.g. corn stalks).
Soil Colloid: smallest particles of the soil system that represents the most active portion of soil that help determine the physical and chemical properties of the soil (e.g. clay particles, humus material).
Soil Management Practices: a broad term to describe any anthropogenic impacts or actions on the soil.
Soil Microbial Community: a broad term used to refer to the diverse grouping of soil bacteria, actinomycetes, fungi, algae and protozoa that interact in a soil ecosystem and their impacts on soil properties.
Synthetic Fertilizers: See inorganic fertilizers.
Substrate: a compound in the soil acted upon by an enzyme to release specific products.
Urea: a nitrogen containing compound that is the end product of the breakdown of protein metabolism. It is often used in synthetic fertilizers as a form of nitrogen.
Urease (UR): an enzyme responsible for converting urea into ammonium carbonate, a compound that readily degrades to ammonia (NH3) and carbon dioxide (CO2).
PDF: Soil Enzyme Interpretation Guide
In all soils, soil microbiology is responsible for creating and releasing soil enzymes to cycle nutrients and to form and decompose organic matter in the soil. The continued promotion of soil health practices (i.e. no-till, diverse cropping systems, organic amendments, etc.) has created an interest in earlier detection of soil management effects through changes in soil microbiological activity. Microbial activity can be reflected in enzyme production, activity and stabilization within the soil. In many cases, soil enzyme activity can respond quickly (i.e. within 2 years) to changes in soil managements when compared to other physical and chemical properties, thus supporting the use of enzyme activity as a “bioindicator” of soil health.
Interpretation of soil enzyme activity requires an understanding of nutrient and organic matter cycling. Often, healthy, active systems have higher enzyme activity, relating to better cycling of nutrients and organic matter quality compared to degraded soils. Below is a list of common soil environmental and soil management characteristics that are often studied. The overlapping impact the following soil factors and soil management strategies have on soil microbial activity causes interpretations to be tied to a singular factor. Thus, this interpretation guide is presented with generalized trends of soil enzyme activity responses to a few soil physical and chemical characteristics as well as soil management practices. This is by no means an exhaustive list as the impact of any desired goal in soil management can be tested.
The soil is often a hostile environment from a microbial perspective. Although enzyme production begins within the cell, extracellular enzymes are exposed to denaturing, predation and adsorption in the soil environment that can inhibit enzyme activity. In addition, the influences of temperature, moisture and pH affects the rate, concentration and structure of an enzyme while the presence of clay and organic matter can provide more adsorption sites and food sources for continued activity for years to come. Increasing water, temperature and soil organic matter will change microbial numbers and community composition while accelerating the activity of enzymes. A few soil physical and chemical characteristics, outlined below, are important to note when understanding and interpreting soil enzyme activity.
Soil organic matter (SOM) is the portion of soil that contains the decomposed products of plants, fauna and microbial biomass. This combination of organic compounds is synthesized by microbial activity and serves as an important source of nutrients and erosion prevention while increasing water holding capacity and soil aggregation. It is also an important habitat for soil microbiology. It is considered a stable portion of soil with only a small portion being mineralized (≈5%) under natural conditions. Soil organic matter mineralization and immobilization is strongly impacted by soil microbial activity. Soil microbial communities can decompose a wide range of plant and animal compounds, allowing accumulation of microbial residues to contribute to SOM formation and stabilization. Exposure of starved microbial communities to newly available food sources in the soil (e.g. incorporating crop residues through tillage) can temporarily increase enzyme activity and continued practice can cause decreases in SOM. This frenzy of activity can be easily noted by the earthy smell of tilled soil. In addition, fluctuation in SOM content due to changes in moisture, temperature and oxygen are also impacted by soil management decisions. In general, higher percentages of SOM support larger microbial populations, and thus contain higher enzyme activity. Enzymes can also become adsorbed on soil humus, the stable portion of SOM, supporting continued activity for several years.
Clay particles consist primarily of microscopic plates that are stacked in a somewhat random manner in the soil. This plate-like structure has a high surface area and net negative charge that greatly influences the physical and chemical properties of soil such as the water and nutrient holding capacity while playing an important role in the adsorption of extracellular enzymes. Once adsorbed on the clay particle surface, the enzyme becomes protected from microbial degradation and environmental stressors. The shape of an enzyme allows for only one active site, or the site where a substrate is acted upon, and depending on how the enzyme is adsorbed to the clay particle, can determine whether the enzyme can act on any further substrates. Although the orientation of the adsorbed enzyme can cause the enzyme to become temporarily inactivated, many of the enzymes will continue contributing to the cycling of soil nutrients for years. The continued activity of adsorbed enzymes supports the use of enzymes as better indicators of long term impacts of soil management decisions. Often, soils with a higher clay content can support more stabilized enzymes and may have higher enzymatic activity.
Soil pH typically reported on a soil test represents the average concentration of hydrogen (H+ ) ions within the soil solution. This measurement, sometimes referred to as bulk pH, can provide valuable information for predicting potential microbial reactions and enzyme activities in soil. Although this measurement is an important overall indicator of the soil pH, localized microsites within the soil can be drastically different than bulk pH readings. Shifts in pH can occur near soil colloids, roots and organic matter in response to biological activity and chemical reactions. Plants and soil microbes can alter the soil environment to free soil nutrients or as a defense mechanism against potential predators or competitors. These pH changes, known as microscale pH, can occur within a few micrometers of these sites but drastically change SOM decomposition, pesticide performance and the solubility and availability of plant nutrients within the surrounding area.
Soil pH is considered a universal regulator of soil enzyme activity. Each soil enzyme has an optimal activity over specific pH values because pH impacts the shape and structure of the enzyme and influences substrate availability. Within this range, the enzyme can effectively act upon the substrate and release nutrients. Beyond this range, changes to the enzyme structure begin to impact productivity. Extreme changes in pH can permanently alter an enzyme’s structure, known as denaturing, and prevents an enzyme from further acting upon any substrates within the soil environment. Soil enzymes within the soil solution are at a higher risk of denaturing from temperature and pH shifts than adsorbed enzymes because adsorbed enzymes adopt a structural adaptation that prevents denaturing. This allows continued enzyme activity even during changes in microscale pH. The pH optimums of adsorbed enzymes can be 1-2 pH units higher than enzymes in soil solution. For example, absorbed soil urease enzymes have a pH optimum of 8.5-9.0, which is about 1-1.5 pH units higher than urease activity in the soil solution. In the laboratory, soil enzyme assays are carried out under pH optimums that produce the highest potential activity for the selected enzyme. The optimum pH for βglucosidase (BG), urease (UR), phosphodiesterase (PHD) are 6.0, 7.5-8.8, and 8.0 respectively. These optimums are based on microsite pH and should not be confused with bulk pH, although drastic changes in bulk pH can strongly impact microsite pH. For ideal conditions, bulk pH should still be in the desired range of pH of 6.0- 7.2 for most crops.
As soil temperatures rise, chemical and enzymatic reaction rates within the soil increase, causing a rapid rate of microbial growth and enzyme production. The production and distribution of enzymes contributes to faster residue decomposition and cycling of nutrients within the soil. Changes in soil temperature are dependent on factors such as climate, season, soil type, plant and residue cover, soil water content and soil organic matter content. Increases in temperature, in addition to moisture, can lead to an increase in soil enzyme production and excretion into the soil environment promoting decomposition and the release of plant available nutrients. Soil temperatures can experience drastic changes seasonally, and even daily. Fortunately, fluctuations in temperatures have caused native soil microbiology to create unique adaptations to survive under harsh conditions. One such adaptation is the ability of microbes to modify the structure of extracellular enzymes to withstand the natural temperature extremes of the ecosystem. Although some soil enzymes may have greater flexibility in the optimal temperature ranges, most soil enzymes still have similar optimal temperature for maximum effective activity. Enzymes active in decomposition (β-glucosidase) and nutrient cycling (urease, phosphodiesterase) are measured in the lab at 98.6°F (37°C).
Seasonal changes in temperature can alter relative rates of decomposition and cause a variation in enzyme activity. Thus, sampling times for soil enzyme activity should be taken when temperature extremes are at a minimum. For most areas, this would be in the fall or spring. Outside these stable temperature periods, shifts in temperatures, moisture availability and substrate availability can impact enzyme activity and cause improper interpretation of enzyme results.
Changes is soil management strategies, soil use, or disturbance are often reflected in the root zone because of the interaction of plant roots and soil microbiology. Plants release readily available carbon-based nutrients to encourage soil microbiology to become established in and around the root zone. The close proximity of plant roots to soil microbiology, and the activity of soil enzymes, allows the release of mobile nutrients to supplement both soil microbiology and the plant via the root. The symbiotic relationship between the plant and microbes creates an active zone within the roots that are sensitive to changes in the soil environment. Enzyme activity tends to decrease with soil depth because root and soil microbe interactions are minimized beyond the root zone. When measuring soil enzyme activity, considerations for tillage depth are important. In soils that experience heavy tillage, soil enzyme samples should be measured to the plow layer depth due to the even distribution of soil nutrients throughout the soil profile. In minimally tilled and no tilled soils, sampling the top 6 inches of soil provides an indication of soil biological activity because residue and nutrient inputs occur near the surface of soil in these systems. When comparing activities between management sites with varying degrees of tillage, make sure all enzyme sampling occurs at the same depth to ensure soil enzyme activities are properly interpreted. Unless there is a specific project goal, we suggest a sampling depth of 6 inches.
Extracellular soil enzymes are dependent on soil moisture to diffuse substrates through the soil environment and become adsorbed to soil surfaces. In a moisture limited environment, soil enzymes can only access substrates that are within a very close vicinity. Depending on the location and concentration of substrates relative to an enzyme, activity of soil enzymes can be strongly impacted. Soils saturated with water for a long period of time may lead to low microbial biomass, mostly attributed to the low oxygen conditions that are unfavorable for aerobic microbes.
Soil management decisions such as tillage management strategies, crop selection and diversity, seed treatments, cropping rotations, and fertilizer managements have compounding and cascading impacts on all aspects of soil. These decisions shape unique soil microbial communities which influence soil properties such as soil structure, porosity, and nutrient availability for plants. The use of soil enzyme activity to compare different soil management strategies can indicate earlier shifts in soil microbial community dynamics and help address management changes before many other soil testing methods. The impacts of commonly used soil management strategies on soil enzyme activities are given below.
Soil structure is formed through physical, chemical and biological activities that create glues to hold soil particles together, called soil aggregation. Different arrangements of aggregates result in different sized pores, creating channels for air and water movement in the soil which ultimately influence bulk density, aeration, permeability and water holding capacity of the soil. Soil aggregation can create areas of high and low oxygen, shield SOM from microbes, and allow diffusion of nutrients and gases through the soil. The mechanical act of tilling a soil damages the aggregate channels and initially flushes the soil with an abundance of air and water, while distributing the once shielded SOM to microorganisms. Most soil microbes are aerobic, or oxygen requiring, organisms that become hyper active in the presence of renewed oxygen and substrates. Invigorated microbial activity leads to greater microbial mineralization and leaching of nutrients from the soil system. Continuous tillage results in poor soil structure, decreased water holding capacity, and lower infiltration rates. In addition, under saturated conditions, microbial activity can consume the limited oxygen causing the areas to become anoxic, or low in oxygen. Lower oxygen levels can inhibit beneficial microbes, allowing pathogenic microbes to thrive. These soil characteristics caused by tillage negatively impact microbial community composition and activity. Furthermore, prolonged tillage practices lead to a loss of aggregate channels and lower substrate availability causing microbes to scavenge nutrients from SOM, resulting in a gradual loss of SOM over time.
In general, soil enzyme activity is found to steadily increase under reduced and no-till fields in comparison to conventional tillage methods. Under no-till systems, application of fertilizers and liming agents are often surface applied and plant residues stay at the surface leading to increases in organic C, N, and P concentrations at the surface soil layer. The presence of increased nutrient concentrations in the soil surface layer promotes increased microbial diversity and activity within the top few inches of soil. In addition, no-till systems or reduced tillage systems allow fungi to thrive. Increased diversity in fungi and bacteria promotes greater activity leading to enhanced aggregation, improved soil structure, stabilized SOM, and increased surfaces for stabilized soil enzymes.
Plants and soil microbes have unique symbiotic relationships in which the plant supplies simple compounds, known collectively as root exudates, to support the colonization of the root by microbial communities. In exchange for the simple compounds, microbes release soil enzymes to mineralize nutrients from the soil for use by the plant and microbe. Individual plants release distinct exudates, encouraging relatively unique microbial communities to exist around the root. Root exudates can shape the soil environment around the root by releasing chemicals that can assist the plant in obtaining nutrients (i.e. iron), avoid pathogens (i.e. allelopathy), and encourage the colonization of beneficial microbes.
Plant diversity, established through a variety of plant types (e.g. perennials, annuals, grasses, broadleaves, legumes, etc.) and root structures (e.g. fibrous, taproot) can present a variety of microhabitats that supports a wider range of microbes. The increased diversity supports greater nutrient competition, prevents the dominance of one or a few microbial species, and generally improves the complexity of the soil microbial community. This soil microbial diversity can be accomplished using cover crops, cropping rotations or a mixture of both. Allowing continuous and diverse vegetative cover mimics a natural soil environment, supporting better nutrient cycling, soil aggregation, soil microbial activity, and water infiltration while suppressing weeds, pathogens and soil erosion. In comparison, a monoculture often reduces microbial and enzymatic diversity because a single crop species creates similar microbial food (e.g. root exudates), habitats (root structures), and chemical signals leading to a dominance of a few organisms in the microbial community. Thus, lower enzyme activity is often found under monoculture systems.
Generally, soil enzyme activity is higher in undisturbed areas than cultivated fields while including cover crops in cultivated fields often increases enzyme activity. Enzyme activity can respond differently to cover crops, indicating the unique soil environment each plant can create. Successful comparisons between cover crop mixes and cropping rotations can assist in future soil management choices.
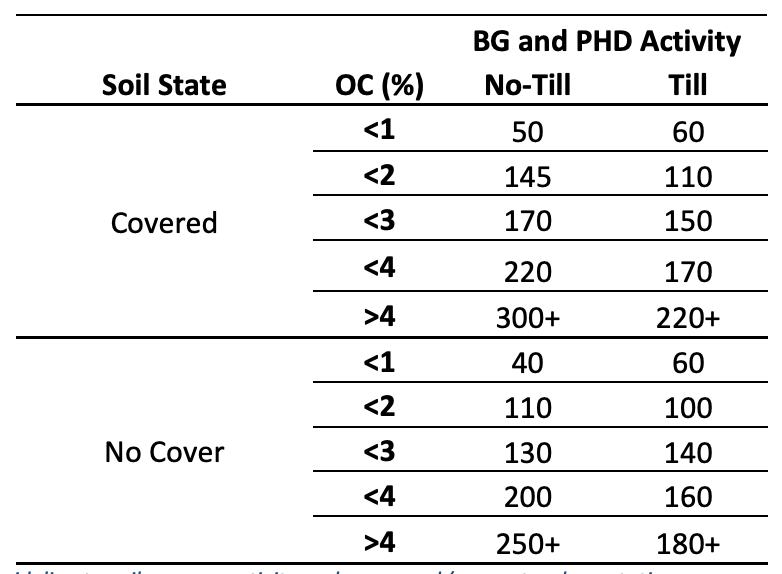
Table 1: A generalized guideline to soil enzyme activity under covered (e.g. natural vegetation, cover crops or diverse cropping rotations) and not covered (e.g. no cover crop, monoculture systems) in different tillage systems and organic carbon (OC) percentages. These are generalized values from literature. Soil enzyme activity is influenced by factors such as those discussed in this interpretation guide and can cause fluctuations in values. Soil enzyme activity should always be used on a comparative basis. Enzyme activity is reported as ppm p-nitrophenyl (pNP) g-1 dw soil h-1 .
Crop production often requires the supplemental input of nutrients as fertilizers for proper growth and desirable yields due to nutrient removal from previous crops. Fertilizers can be subdivided into organic (i.e. manure, compost) and inorganic (i.e. chemical fertilizer) categories. Fertilizer type, composition and concentration can impact soil community dynamics and enzyme activity. Generally, organic amendments contain carbon-based complex compounds that require the activity of soil microbiology mineralization to convert the nutrients into plant available forms. Inorganic fertilizers primarily exist in plant available forms, requiring little to no microbial conversion. Synthetic fertilizers may also be consumed by the microbes, leading to a temporary increase in soil microbial community dynamics and activity. However, some inorganic fertilizers can inhibit enzyme production. For example, ammonia-based N fertilizers are often treated with a urease inhibitor that can decrease urease activity with increasing application.
The application of fertilizers (especially N based fertilizers) can have varied impacts on enzymes dependent on application rate, amount, vegetation, and other management decisions. Generally, higher enzyme activity is found under organic fertilization as opposed to inorganic fertilization. Soil microbial communities respond quickly to fertilizer inputs which may skew data if sampled shortly after soil fertilizer application. Sample at least 2-3 months after soil amendments such as manure or fertilizer application.
The Haney, phospholipid fatty acids (PLFA), and enzyme tests depict a holistic understanding of nutrient availability and dynamics, microbial community abundance and diversity, and enzymatic response to changing soil environmental conditions.
The Haney test utilizes a combination of soil testing methods: soil respiration, water extract and H3A extract. Soil respiration indicates a measurement of the soil’s microbial biomass and potential for activity under optimal conditions. This test quantifies the amount of CO2-C a soil can produce over a 24-hour incubation period following a significant drying and rewetting event. The water extract measures the amount of soluble organic carbon and nitrogen readily available to microbes in the soil. The H3A extract contains organic acids that mimic root exudates and provides a measurement of the nutrients the plant will be able to use during the growing season.
The Phospholipid Fatty Acid (PLFA) test provides an analysis of the total living microbial biomass in soil. This analysis depicts the abundance of bacteria, actinomycetes, rhizobia, arbuscular mycorrhizal fungi, saprophytic fungi and protozoa by quantifying specific fatty acids known as biomarkers. Also included are the fungi:bacteria ratio, predator: prey ratio, and gram positive: gram negative bacteria ratio. The PLFA test is a useful tool to monitor microbial community response over a number of years or across different land management practices.
The Haney Test, PLFA and enzyme tests can all be combined to display a unique look a soil from a microbial perspective. Soil microbes produce enzymes based on the availability of substrates or lack of nutrients within the soil environment and is strongly correlated with many of the organic nutrients in the soil. Changes in soil enzyme activity can provide early indications of shifts in microbial nutrients in the soil, as measured by the Haney test, and shifts in the microbial community structure and abundance, as measured by the PLFA. For instance, βglucosidase activity strongly correlates with organic carbon content in soil and can indicate early changes in SOM. The use of all three tests can provide a wealth of information on the soil microbial environment.
Soil microbiology produce and exude soil enzymes to cycle nutrients (e.g. urease, phosphodiesterase) and energy (e.g. β-glucosidase) in the soil. Because enzymes are nutrient specific, monitoring fluctuations in potential enzyme activity can be a useful tool in gauging the microbial response to changes in soil managements. Soil enzyme activity comparisons between land management strategies or over successive years may provide earlier indications of SOM generation and destruction and nutrient availability. Changes in key enzyme activities can signal a higher nutrient input and cycling in the soil that can lead to building healthier soils. Healthier soils can aid in weed suppression, lower dependence on fertilizers and store greater amounts of nutrients. This soil enzyme interpretation guide is intended as a resource to aid producers in understanding soil enzyme activity and the role it plays in building healthier soils. To learn more about the role soil enzymes play in the soil or about specific soil enzymes, please visit our website. This interpretation is by no means limited to the above soil environment and management factors as soil enzyme activity may be used to detect the impact of any desired soil management strategy.
Additional information will be added to the website as new information becomes available. Any questions regarding soil enzyme testing may be directed to Ward Laboratories, Inc at biotesting@wardlab.com.
PDF: Specific Soil Enzyme Information: Betaglucosidase Test
Energy is harnessed in a plant through photosynthesis, a process by which carbon dioxide (CO2) and water (H2O), with the help of energy from the sun, is converted to different carbohydrates and oxygen (O2). Plants store and use the various forms of carbohydrates, such as sugars, starches and cellulose, for maintenance, cell structure and growth. In general, carbohydrates are compounds that contain different ratios of carbon and water molecules that serve as a fundamental source of energy and that emphasizes the importance of the carbon cycle in the soil. In addition, plants can release carbohydrates from the roots to help cultivate distinct soil microbial communities, which use the carbohydrates as a main source of chemical energy.
When a plant dies, soil microbiology begins decomposing the strong, rigid structure of cellulose, the basic structural component of plant cell walls and fiber. Soil microbes produce a collective group of soil enzymes, known as cellulases, to disassemble the rigid structure of cellulose to release glucose molecules, a simple readily available sugar and important source of easily accessible energy for soil microbes. The cellulose structure is fractured by three dominant groups of enzymes: endoglucanases, exoglucanases and β-glucosidases (See Figure 1). Endoglucanases randomly cleave the long-chained cellulose into smaller fractions while exoglucanases further reduce the cellulose chain to cellulobiose units (two linked glucose molecules). Finally, β-glucosidases cleave the cellulobiose molecules to glucose, a readily available energy source for plant and microbial uptake.
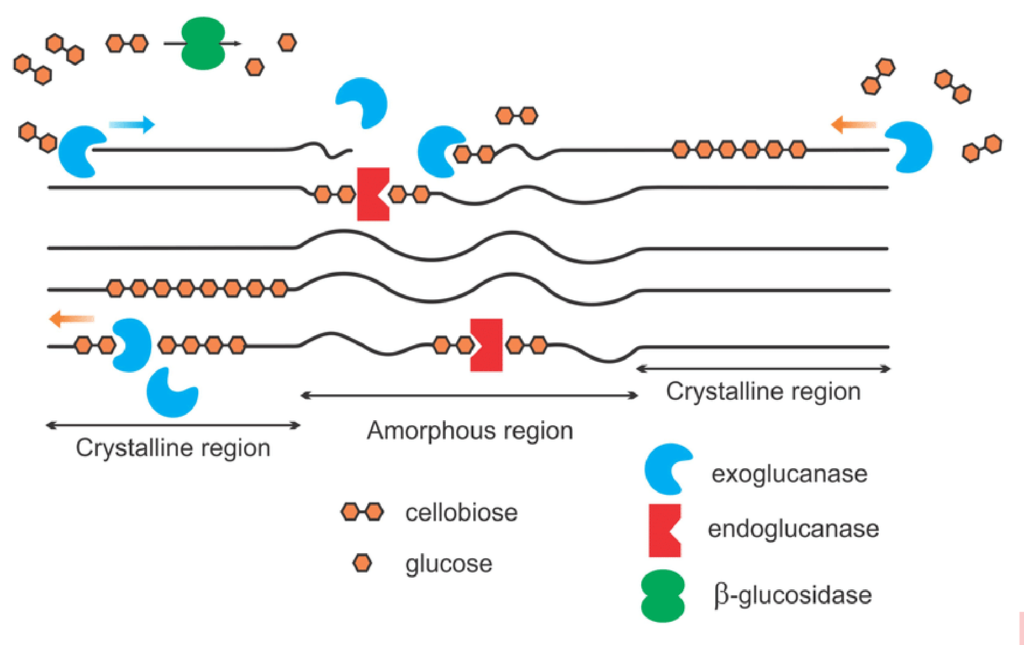
Figure 1: Simplified schematic of cellulase enzymes acting on cellulose
The long-chained cellulose is fractured by the accumulated effort of exoglucanases, endoglucanases and β- glucosidase enzymes to release glucose in the soil environment. (Image from Akhtar 2016)
Soil organic matter (SOM) serves as an important habitat for soil organisms and is a source of short- and long-term carbon storage. Increasing SOM provides numerous benefits to the soil such as improved soil structure and water retention. Traditional SOM measurements are based on a percentage of the whole soil system, making small, subtle changes in SOM due to changes in soil managements difficult to detect. Soil organic carbon (SOC) represents carbon stored in soil organic matter and the quantity and quality of SOC can indicate the availability of energy and nutrients for plants and soil microbes. Because of the pivotal role β-glucosidase (BG) plays in residue decompositions and the carbon cycle, the activity of BG is closely monitored to detect changes in the carbon cycle and SOC cycling in a shorter time span (approximately 2 years compared to 5 – 7 years through traditional laboratory methods). Greater soil microbial biomass often has higher BG activity indicating a soil microbial community’s ability to break down plant residues and cycle nutrients through the soil, an important step to ensure nutrients are available in the soil for future crops. Thus, BG can provide an early indication of changes in SOC sooner than traditional total and organic C analysis can discern differences. Absence or suppression of BG prevents or reduces the cycling of soil nutrients that can impact plant health and alert producers to early issues in soil health.
Soil enzyme testing at Ward Laboratories is conducted by analyzing the consumption of a substrate and the release of a colored product. The consumption of product is measured over time and results are expressed as a rate of enzyme activity. By using controlled soil conditions (e.g. pH, temperature), the enzyme activity rates can indicate the potential activity for the soil enzymes under ideal conditions. This allows a comparison of potential enzyme activities between different soil management practices. Similarly, the same site can be tracked over time to monitor subtle changes in microbial dynamics and provide an indication of the microbial community response to changing environmental conditions and management.
Soil samples received by the laboratory should be cooled and in field moist condition. Each soil sample is passed through a 2 mm sieve and weighed into two centrifuge tubes (1.00 ± 0.05 g each). The first subsample is referred to as the treatment sample and the second subsample is referred to as the control sample. Each vial receives 4 mL buffer solution. The treatment vial receives 1 mL substrate prior to all samples being incubated for 1 h at 37°C. After incubation, the control vial receives 1 mL substrate. All vials receive 4 mL stop buffer and 2 mL flocculant. Vials are centrifuged, filtered with Whatman 2V filter paper and analyzed on a spectrophotometer at 405 nm. Enzyme activity is expressed as ppm p-nitrophenol g-1 dry weight soil h-1.
Interpretation of soil enzyme activity requires an understanding of nutrient or organic matter cycling. Often, healthy, active systems have increased enzyme activity, relating to better cycling of nutrients and organic matter quality in the soil. Nevertheless, sites with recent disturbances may have higher activity levels due to increased substrate availability when compared to non- disturbed sites. For example, a conventional tillage field versus a no-till field in its first year of transition may indicate the tilled field has higher enzyme activity. This is because the act of tilling a field provides aeration and better distribution of substrates to microbes. The increased activity cannot be fully sustained in this system and often causes the enzymes to access the nutrients in organic matter, leading to a loss of organic matter the following year. A list of common soil characteristics and soil management impacts can also be found in the interpretation guide.
Additional information will be added to the website as new information becomes available. Any questions regarding soil enzyme testing may be directed to Ward Laboratories, Inc at biotesting@wardlab.com.
PDF: Specific Soil Enzyme Information: Phosphodiesterase Test
Plant available phosphorus (P) is dependent on the immobilization, adsorption and dissolution of P in the soil environment. As the second most limiting nutrient, P is often applied in standard fertilizer applications. Adequate P availability is important to support cell growth and stimulate early plant growth and maturity in crops. The mineralization and cycling of organic P in the soil is dependent on the activity of specific soil enzymes, known collectively as phosphatases, to transform organic P to plant available P. Without the activity of phosphatases, a substantial proportion of P would be immobile or structurally unavailable to plants and soil microbes. Phosphatases act on organic P within the soil in addition to commonly applied fertilizers, such as manure, to form free phosphates (PO43-) available for plants and soil microbiology.
Phosphodiesterase (PHD) is a phosphatase responsible for the degradation of nucleic acids, phospholipids and other diesters in the soil. When fresh organic P enters the soil, phosphodiesterase acts as the initial responder and cleaves the large compounds into smaller, easily accessible molecules (See Figure 1). Depending on the soil pH, alkaline or acid phosphomonoesterase enzymes further reduce the molecule to free phosphates. Thus, PHD activity is tightly linked to soil organic P cycling in the soil. In addition, PHD activity is often higher in the rhizosphere than in bulk soil, suggesting a significant role of phosphodiesterase in plant P acquisition.
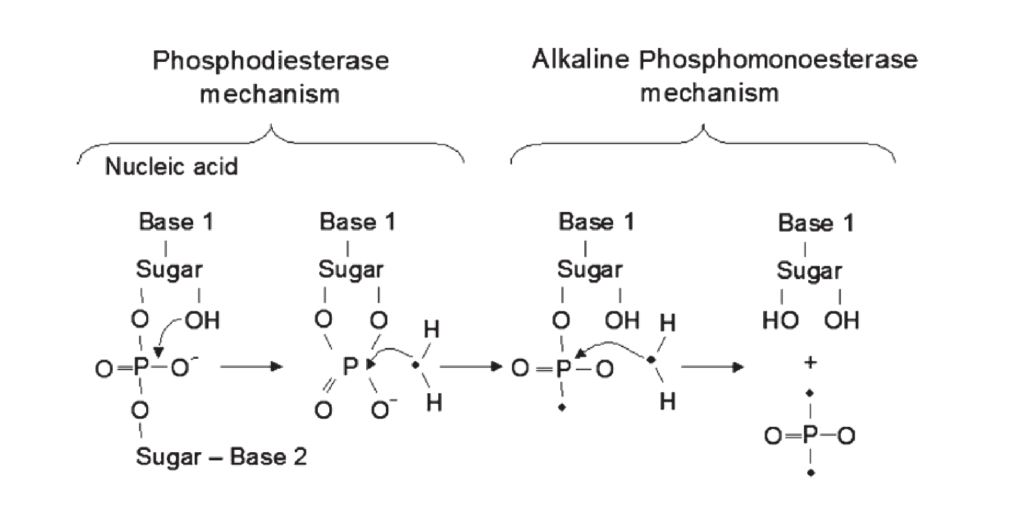
Figure 1: The action of phophodiesterase and alkaline phosphomonoesterase to release phosphates
(Image from Nannipieri 2011)
Soil enzyme testing at Ward Laboratories is conducted by analyzing the consumption of a substrate and the release of a colored product. The consumption of product is measured over time and results are expressed as a rate of enzyme activity. By using controlled soil conditions (e.g. pH, temperature), the enzyme activity rates can indicate the potential activity for the soil enzymes under ideal conditions. This allows a comparison of potential enzyme activities between different soil management practices. Similarly, the same site can be tracked over time to monitor subtle changes in microbial dynamics and provide an indication of the microbial community response to changing environmental conditions and management.
Soil samples received by the laboratory should be cooled and in field moist condition. Each soil sample is passed through a 2 mm sieve and weighed into two centrifuge tubes (1.00 ± 0.05 g each). The first subsample is referred to as the treatment sample and the second subsample is referred to as the control sample. Each vial receives 4 mL buffer solution. The treatment vial receives 1 mL substrate prior to all samples being incubated for 1 h at 37°C. After incubation, the control vial receives 1 mL substrate. All vials receive 4 mL stop buffer and 2 mL flocculant. Vials are centrifuged, filtered with Whatman 2V filter paper and analyzed on a spectrophotometer at 405 nm. Enzyme activity is expressed as ppm p-nitrophenol g-1 dry weight soil h-1.
Interpretation of soil enzyme activity requires an understanding of nutrient or organic matter cycling. Often, healthy, active systems have increased enzyme activity, relating to better cycling of nutrients and organic matter quality in the soil. Nevertheless, sites with recent disturbances may have higher activity levels due to increased substrate availability when compared to non- disturbed sites. For example, a conventional tillage field versus a no-till field in its first year of transition may indicate the tilled field has higher enzyme activity. This is because the act of tilling a field provides aeration and better distribution of substrates to microbes. The increased activity cannot be fully sustained in this system and often causes the enzymes to access the nutrients in organic matter, leading to a loss of organic matter the following year. A list of common soil characteristics and soil management impacts can also be found in the interpretation guide.
Additional information will be added to the website as new information becomes available. Any questions regarding soil enzyme testing may be directed to Ward Laboratories, Inc at biotesting@wardlab.com.
